Artificial intelligence in breast cancer diagnosis through histopathology and biomarker detection: a scoping review
- Faculty of Health Sciences, Universiti Teknologi MARA, Cawangan Pulau Pinang, Kampus Bertam, 13200 Pulau Pinang, Malaysia
- Department of Biomedical Sciences, Advanced Medical and Dental Institute (AMDI), Universiti Sains Malaysia, Kepala Batas Penang, 13200 Malaysia
Abstract
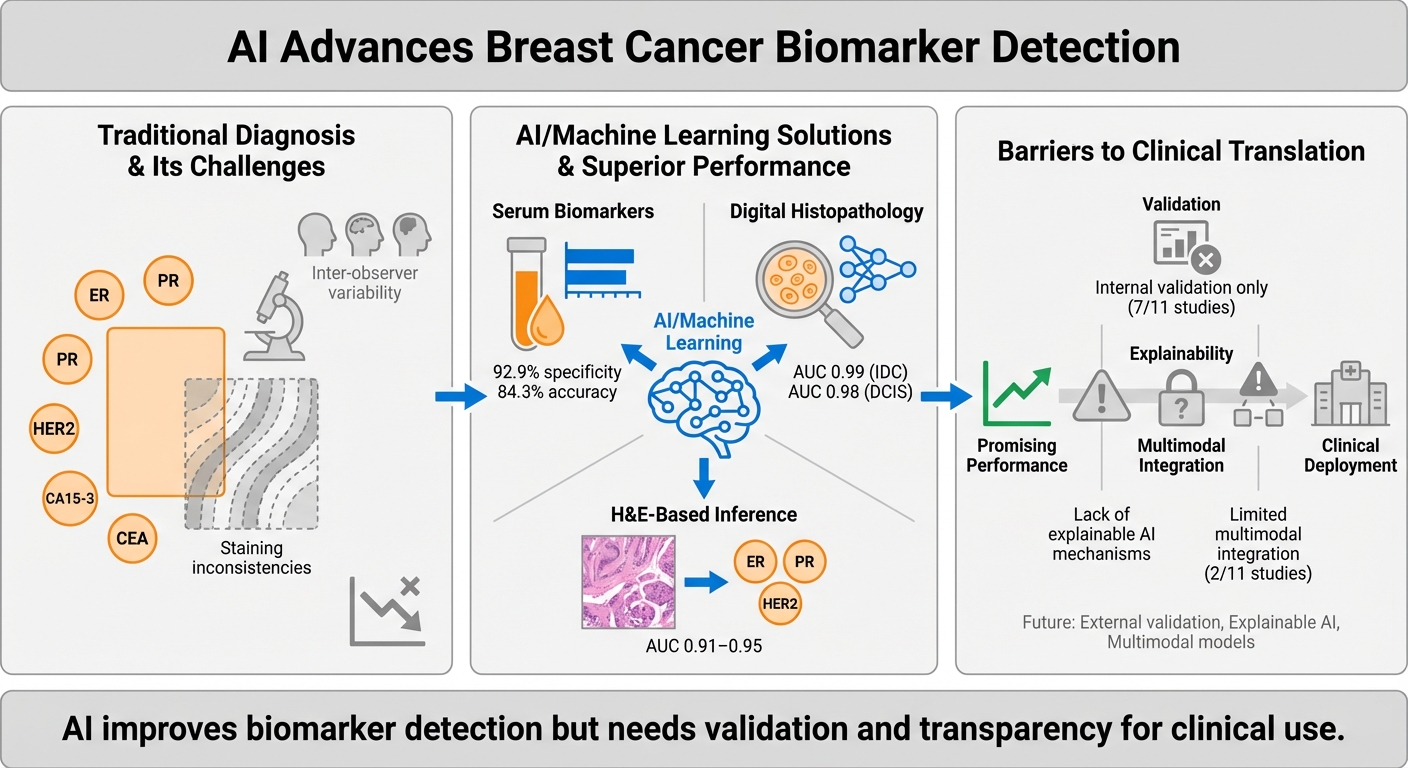
Breast cancer diagnostics depend critically on the accurate histopathological evaluation and identification of tumour biomarkers in biopsies and serum. While standard operating procedures (SOPs) provide structured diagnostic workflows, they are constrained by inter-observer variability, staining inconsistencies, and inconsistencies in interpretive criteria and decision thresholds. Artificial intelligence (AI) offers promising solutions by integrating heterogeneous data sources and standardising interpretation. This scoping review critically evaluates 11 recent studies on AI-driven approaches for tumour biomarker detection using serum panels, immunohistochemistry (IHC) images, digital histopathology, and morphology-based prediction. AI techniques such as convolutional neural networks, gradient-boosted machines, and ensemble models, have demonstrated improvements in diagnostic performance, reproducibility, and efficiency, achieving up to 92.9% specificity in serum-based models and image-based classifications. Several studies have shown that AI can infer molecular biomarker status directly from H&E-stained slides, thereby eliminating the need for biochemical assays. Despite these advances, significant challenges persist. Many models lack external validation, explainability, and integration into clinical workflows. Furthermore, concerns regarding data bias, ownership, and equitable deployment highlight the urgent need for ethical governance and inclusive model design. Future research should prioritise prospective validation, multimodal data integration, and clinician-centred implementation strategies to realise AI’s potential in advancing breast cancer diagnostics and personalised care.
Introduction
Tumour biomarkers are measurable biological indicators that capture the molecular and pathological features of breast cancer, and they play a central role in diagnosis, prognosis, and treatment planning. Clinically relevant biomarkers include hormone receptors such as estrogen receptor (ER), progesterone receptor (PR), and androgen receptor (AR), serum markers such as CA 15-3, CA 27.29, carcinoembryonic antigen (CEA), and nestin, and molecular indicators such as human epidermal growth factor receptor 2 (HER2), antigen Kiel 67 (Ki-67), and breast cancer gene 1 and 2 (BRCA1/2). These markers are routinely assessed through IHC, genetic testing, and blood-based assays, with their results guiding therapeutic decisions and enabling targeted interventions1.
Despite standard operating procedures (SOPs) intended to ensure reproducibility, several clinical and technical challenges affecting biomarker reliability have arisen from these procedures. Sources of variability include differences in tissue handling, fixation, staining protocols, and subjective interpretation of borderline cases2,3. Serum biomarkers are further limited by low sensitivity and specificity in early-stage disease4. More importantly, current diagnostic frameworks often treat histological, molecular, and serum data in isolation, overlooking their interplay in shaping tumour behaviour and therapeutic response1,5. This fragmented approach risks inconsistent diagnosis, delayed treatment, and reduced capacity for personalised care.
AI offers a potential solution by integrating multimodal data of histopathology, IHC, genomic, and serum profiles into unified, interpretable outputs. Deep learning models have already demonstrated accuracy comparable to human experts in breast cancer subtyping, biomarker prediction and recurrence risk assessment6,7. However, challenges of explainability, validation, and workflow integration remain. Against this backdrop, the present scoping review systematically maps recent applications of AI in tumour biomarker detection, with a focus on histopathology, IHC, serum-based assays, and morphology-to-biomarker inference. Unlike prior reviews that emphasised AI in image-based diagnosis, this synthesis adopts a biomarker-focused perspective and incorporates the most recent contributions up to June 2025, such as in Garberis et al. (2025) and Tanveer et al. (2025). This narrower, biomarker-focused lens distinguishes our synthesis from prior reviews and highlights methodological gaps directly relevant to personalised breast cancer diagnostics.
Methods
We performed a scoping review in accordance with the PRISMA-ScR guidelines. The following search string was used: (“breast cancer”) AND (“tumor biomarker” OR “biomarker”) AND (“artificial intelligence” OR “machine learning” OR “deep learning”) AND (“histopathology” OR “histology” OR “hematoxylin eosin”). The search was limited to English-language articles published between January 2018 and 30 June 2025. We identified PubMed (n=125), Scopus (n=105), Web of Science (n=97), additional records were identified through manual reference screening (n=15). The full search syntax per database is shown in Supplementary 1. Duplicates were removed prior to screening, so per-database exclusion counts are not reported separately.
A scoping-review design was selected to comprehensively map the breadth and methodological characteristics of AI applications in tumour-marker detection across multiple diagnostic modalities (serum biomarkers, immunohistochemistry, histopathology, and molecular assays). This approach was appropriate given the heterogeneity of study designs and outcomes, which precluded meta-analysis.
In addition to peer-reviewed journal articles, grey literature (including conference proceedings and preprints indexed in the above databases) was screened for relevance. Two reviewers independently conducted the search and screening process, resolving disagreements through discussion and consensus. This scoping review was conducted in accordance with the PRISMA-ScR guidelines. No preregistration was performed and no professional librarian was involved. However, the two-reviewer system enhanced methodological rigour and reproducibility.
Although scoping reviews primarily aim to map evidence rather than exclude studies by quality, a simplified critical appraisal was undertaken to contextualise methodological robustness. Each included study was assessed using the CHARMS-SF framework for prediction and diagnostic modelling, which covers study design, data source representativeness, sample size, predictor handling, outcome definition, model development, validation strategy, and reporting completeness, as described in Moons et al. (2014)8. Two reviewers independently performed the appraisal and resolved discrepancies by consensus. The objective was not to exclude studies but to transparently identify potential sources of bias and applicability limitations (Supplementary 3 and 4).
All authors contributed to the conceptualisation, design, and writing of the manuscript. During manuscript preparation, ChatGPT and QuillBot were used exclusively for language refinement, rephrasing, grammar correction, and enhancing narrative coherence. No new scientific content was generated, and all sections involving data extraction, analysis, or interpretation were written and verified by the authors. Outputs from generative AI were fully reviewed for accuracy, fidelity to meaning, and compliance with scientific standards. The authors retain full responsibility for all scientific content.
Results
A total of 327 records were identified from PubMed, Scopus, and Web of Science, with 278 removed as duplicates before screening. Forty-nine were deemed eligible based on the abstract. Of these, 40 were excluded for specific reasons: 12 did not involve any AI model or algorithm, 18 were not focused on breast cancer or tumour biomarkers, and 10 were non-peer-reviewed works such as conference abstracts. Manual reference checks identified an additional 15 records, of which 13 were irrelevant or duplicates. Finally, 9 studies retrieved through database searches and 2 from manual reference checks made a total of 11 studies included in this review. The PRISMA-ScR flow diagram (Figure 1) and Supplementary 2 provide specific reasons for exclusion.
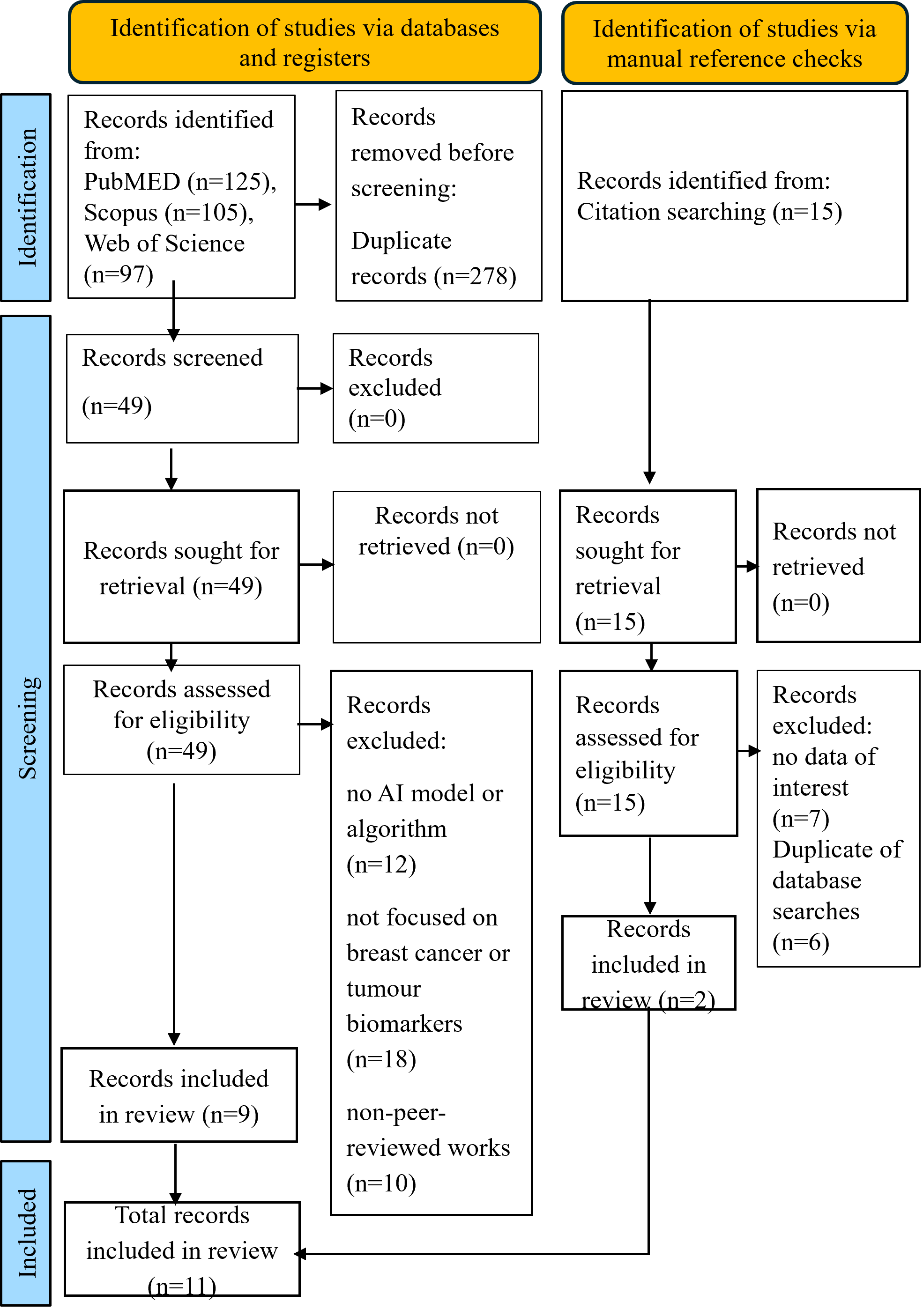

The PRISMA-ScR flow diagram illustrating the identification, screening, eligibility assessment, and final inclusion of studies in this scoping review (n=11). The database search was performed according to the PRISMA 2020 protocol.
Although over 500 AI-related breast-cancer studies have been indexed since 2018, only 11 met our inclusion criteria, as we limited the selection to AI models directly applied to tumour-marker detection or biomarker-driven diagnosis, excluding purely radiologic or survival-prediction pipelines. This narrow focus ensures alignment with the review’s aim of evaluating the integration of AI into current biomarker SOPs.
Our simplified CHARMS-SF appraisal (Supplementary 3), aligned with domains from PROBAST and QUADAS-AI, indicated that most included studies were retrospective, relied on internal validation, and were constrained by small sample sizes, particularly in Dodington et al., or by single-centre or public datasets such as The Cancer Genome Atlas (TCGA) without external testing. Standard performance metrics were inconsistently reported; several studies provided only area under the curve (AUC) values, omitting sensitivity, specificity, or calibration analyses, and few described model interpretability or clinical deployment strategies. Where available, diagnostic performance was expressed as AUC values; however, none of the included studies provided complete 2×2 diagnostic data (true/false positives and negatives) to enable full reconstruction of sensitivity, specificity, or predictive values. Calibration and decision-curve analyses were absent across all studies, limiting assessment of clinical utility and model reliability. Quantitative synthesis confirmed these observations: seven of eleven studies (7/11) used only internal validation, five (5/11) involved fewer than 100 samples, and nine (9/11) were retrospective. Only four (4/11) incorporated any form of external or multicentre validation, and three (3/11) implemented explainable AI (XAI) approaches such as saliency or attention mapping. None reported calibration or decision-curve analyses. Collectively, these findings suggest that although technical performance is encouraging, the current evidence remains preliminary and should be interpreted cautiously, pending broader external validation and prospective impact studies (Supplementary 4).
Studies were categorised into three key diagnostic domains based on their primary AI application: (1) detection of tumour biomarkers in serum, (2) digital histopathology, and (3) prediction of tumour-biomarker expression and treatment outcome in breast cancer patients using histopathology data. The key findings in each study were outlined in Table 1. A narrative synthesis summarised these findings, supported by a visual schematic that highlighted the scope, limitations, and future directions of the reviewed AI models.
Summary of key findings in selected reviewed studies based on primary AI applications: (1) detection of tumour biomarkers in serum, (2) digital histopathology, and (3) prediction of tumour-biomarker expression and treatment outcome in breast-cancer patients using histopathology data. AI = Artificial Intelligence; ML = Machine Learning; GLM = Generalised Linear Model; GBM = Gradient Boosting Machine; RF = Random Forest; CNN = Convolutional Neural Network; XAI = Explainable Artificial Intelligence; H&E = Hematoxylin and Eosin stain; HES = Hematoxylin Eosin Saffron stain; ER = Estrogen Receptor; PR = Progesterone Receptor; HER2 = Human Epidermal Growth Factor Receptor 2; IHC = Immunohistochemistry; TCGA = The Cancer Genome Atlas; IDC = Invasive Ductal Carcinoma; DCIS = Ductal Carcinoma In Situ; FC layers: Fully Connected layers (in neural networks).
| Study | AI technique/ model | Focus area | Conventional SOP assisted | Dataset and cohort size | Study design | External validation | Performance metrics | Code availability | Notes |
|---|---|---|---|---|---|---|---|---|---|
| (Garberis et al., 2025) | RlapsRisk (deep learning, self-supervised, multiple instance learning) | Predict 5-year metastasis-free survival (MFS) from HES-stained slides in ER+/HER2– early breast cancer | Hematoxylin eosin saffron (HES) | GrandTMA (n=1,429, retrospective), CANTO (n=889, prospective) | Development 🡪retrospective training 🡪 prospective external validation | Yes. Validated in prospective CANTO cohort | C-index improved from 0.76 to 0.80 with AI. Significant prognostic value beyond clinico-pathological models | No -proprietary pipeline | First large-scale AI prognostic model externally validated in a prospective trial. Interpretable via Shapley tiles |
| (Mao et al., 2023) | Ensemble ML, OncoSeek platform (GLM, GBM, RF) | Multi-cancer early detection using 7 serum protein biomarkers (CEA, CA15-3, CA125, CA19-9, AFP, CYFRA21-1, NSE) | Serum tumour biomarker for breast cancer (CEA, CA15-3) | Case-control study. 3 cohorts (n~10,000) | Retrospective case-control | Not specified | Specificity 92.9%, accuracy 84.3% | No | OncoSeek used for the first time |
| (Sandbank et al., 2022) | Deep CNN (Ibex Galen Breast, commercial) | Breast biopsy based on grading: invasive, DCIS, atypia, benign lesions | Immunohistochemistry (IHC), hematoxylin and eosin (H&E), hematoxylin eosin saffron (HES) | Training/test: 2,252 slides, 1,090 patients; external: 841 WSIs, 436 patients | Development 🡪 internal test 🡪 blinded external validation 🡪 clinical deployment | Yes. Blinded, multi-site + real-world deployment | AUC 0.99 (IDC), 0.98 (DCIS). Sensitivity 95%, specificity 93% | No – proprietary (Ibex) | Deployed clinically as second-read system |
| (Meshoul et al., 2022) | Feature selection + ML (SVM, RF, Extra Trees, XGBoost); SHAP for explainability | Breast cancer molecular subtype classification, ER/PR/HER2 prediction (TCGA-BRCA omics) | H&E as IHC alternative | TCGA-BRCA and local H&E sets (~1,100 cases) | Computational methods study | No. Internal CV only | Best accuracy with XGBoost & Extra Trees; SHAP provided feature interpretability. AUC > 0.89 | No | Novel integration method, but no external validation |
| (Gamble et al., 2021) | CNN | Predict ER, PR, HER2 biomarker status from H&E slides | Detection of ER, PR, HER2 | TCGA-BRCA (annotated H&E with matched IHC labels) | Retrospective, single dataset | No. Internal only | AUC: ER 0.91, PR 0.95, HER2 0.93 | No | Strong discriminative power, but no external validation, no sensitivity/specificity |
| (Huang et al., 2023) | CNN with cross-domain learning | Robust breast cancer detection across H&E staining protocols | H&E + staining variations | Multi-stain datasets; sample size not fully detailed, multi-institution | Retrospective multi-institution | Yes. Tested on cross-staining datasets | Accuracy >90% across stains; robust to lab/stain variability | No | Shows stain-robustness, but no feature attribution |
| (Luan et al., 2023) | Same as Mao (OncoSeek platform) | Multi-cancer early detection, serum markers | Serum tumour biomarker (CEA, CA15-3) | Multi-centre, China: ~11,000 participants (cases + controls) | Large-scale, multi-centre, retrospective | Yes. External, multi-site in China (urban + rural) | Accuracy 84.3%, specificity 92.9% | No | Independent validation of Mao et al., robust across sites |
| (Dodington et al., 2021) | Supervised ML (nuclear morphometrics) | Predict pathological complete response (pCR) to NAC in high-risk breast cancer | IHC, H&E | n = 82 patients (digitised H&E biopsies) | Retrospective | No. Single-centre only | F1 = 0.87 (outperformed logistic regression baseline) | No | Small sample, no external validation; showed morphometry predicts pCR |
| (Malherbe, 2021) | Narrative/conceptual with computational perspective | Role of AI in tumor microenvironment modeling | Multiplex IHC, spatial omics | Review perspective (not empirical study) | N/A | N/A | N/A | N/A | Highlights limitations of CNNs, proposes GNNs and spatial transcriptomics |
| (Tanveer et al., 2025) | Ensemble ML (radiomics + serum protein markers, meta-classifier) | Early breast cancer detection | Radiomics + tumour biomarkers | Dataset not clearly reported; retrospective | Retrospective | No | Accuracy 91.2% | No | Early detection using multi-diagnostic sources |
| (Jones et al., 2022) | Hybrid ML+DL pipeline (CNN + FC layers) | Integrates mammography + clinical metadata to predict malignancy and molecular subtype | Mammography and H&E | Imaging data (mammography and H&E slides) and electronic health records. n not specified | Retrospective | No |
Reported improved prediction vs. baselines. AUC, sensitivity, and specificity were not measured | No | Lacks interpretability and data governance discussion |
Discussion
AI use in breast cancer diagnosis through the detection of tumour biomarkers in serum
The study by Mao et al. (2023) introduces OncoSeek, an AI-driven model trained on seven serum protein tumour biomarkers, including CEA and CA15-3, for multi-cancer early detection9. The conventional method, which relies on individual thresholds for tumour biomarkers, produced high false-positive rates with a specificity of 56.9%. This means that when fixed cutoff values are used to interpret markers like CEA or CA15-3, many people without cancer are incorrectly identified as having the disease. This high false-positive rate highlights the limitations of conventional threshold-based diagnostics and sets the stage for AI models that can interpret complex patterns across multiple markers. OncoSeek significantly improves specificity to 92.9% and accuracy to 84.3%. This is achieved by employing machine learning models such as generalised linear models (GLM), gradient-boosted machines (GBM), and random forests (RF). These models are capable of capturing non-linear marker interactions. Such interactions involve complex, non-additive relationships among biomarkers, where the effect of one marker on diagnosis depends on the presence or concentration of others. This approach offers a significant advantage over fixed cutoff values, which treat each biomarker independently and fail to account for their interdependencies, often resulting in higher rates of false positives or false negatives. Unlike linear models that assume fixed individual contributions, AI models such as GBM and RF can capture these intricate dependencies, therefore improving diagnostic sensitivity and specificity by learning synergistic or conditional patterns across multiple biomarkers9. This demonstrates AI’s potential to aggregate weak individual signals from multiple markers into a clinically robust diagnostic output.
Luan et al. (2023) confirmed the findings of Mao et al. by demonstrating that tumour biomarkers detection assisted by AI can achieve high specificity in multi-cancer early detection. The same OncoSeek platform was used, which processed inputs from seven serum-based protein tumour biomarkers including CEA and CA15-3, which are specific to breast cancer. The remaining 5 markers AFP, CA125, CA19-9, CYFRA21-1, and NSE are associated with other cancers, enabling the model to support multi-cancer detection10. Each patient’s serum profile was transformed into a feature vector containing marker concentrations, which were labelled based on their confirmed cancer status and tissue of origin. During training, the model learned to assign probabilities to multiple cancer types simultaneously, optimising its performance using stratified cross-validation and grid search. Importantly, this approach enabled the model to detect synergistic patterns between markers. For example, elevated CA15-3 was interpreted as more significant when CEA or other markers were concurrently abnormal. The end result was a marked increase in diagnostic accuracy of up to 84.3% and specificity of 92.9%, particularly in distinguishing malignant from benign or healthy cases, compared to fixed-threshold rules10. While Mao et al. (2023) introduced and validated OncoSeek’s diagnostic performance in a large population, Luan et al. (2023) conducted an independent evaluation of the same OncoSeek platform across multiple institutions in China, involving both urban and rural clinical centres. This multi-site validation confirmed that OncoSeek maintained high diagnostic performance with specificity of 92.9% and accuracy of 84.3%, even when tested on diverse patient populations and laboratory environments9,10. The consistency of results across institutions demonstrated the model’s robustness, highlighting its potential for widespread clinical adoption and reliable integration into real-world workflows regardless of institutional variability. Both studies employed the same AI model architecture and seven-marker serum panel, reinforcing the robustness and reproducibility of OncoSeek for multi-cancer early detection.
AI use in breast cancer diagnosis through histopathology
Huang demonstrated that AI models can maintain consistent diagnostic performance even when applied to histological slides stained using different protocols11. This work lays the foundation for evaluating AI resilience across diverse real-world laboratory settings. These stains were primarily H&E and its colour-normalised variants, excluding antigen-based methods such as IHC or immunofluorescence. Their study focused on developing a convolutional neural network (CNN)-based model that is robust to variations in routine H&E staining caused by differences in reagents, laboratory protocols, and microscopes. By employing cross-domain learning strategies and training on multi-stain datasets, the model achieved over 90% classification accuracy in distinguishing between benign and malignant lesions. It was also able to differentiate among histologic subtypes with high accuracy. These results were observed in multi-institution validation sets that included cross-stained H&E variants11. This increases the model’s robustness to real-world staining inconsistencies. However, the study did not explore the model’s interpretability or offer insights into which morphological features were most predictive. This limits its potential utility as a decision support tool for pathologists. Additionally, while classification accuracy was high, the study did not report sensitivity and specificity for each individual staining domain. Without these detailed metrics, it is challenging to identify specific scenarios where the model may fail (failure modes) or determine how reliably it performs across different histologic classes (class-specific reliability). Nonetheless, the model demonstrated strong cross-stain robustness, which supports its generalisability. This capability offers promise for deployment in real-world laboratories where staining inconsistencies are unavoidable. In summary, Huang’s study illustrates a feasible pathway for integrating AI into pathology workflows that rely on H&E staining.
In the work by Sandbank, a CNN-based model was trained to analyse whole slide images (WSIs) of breast biopsies stained with H&E and hematoxylin-eosin-saffron stain (HES), which adds saffron to enhance the contrast of connective tissue12. The algorithm learned to classify 51 morphological and diagnostic classes, including ER/PR/HER2-related subtypes, various grades and ductal carcinoma in situ (DCIS). Training was performed on a dataset of digitised WSIs, with expert-annotated regions of interest. Through supervised learning, the CNN extracted features such as nuclear morphology, tissue architecture, and staining intensity. Model optimisation was performed using backpropagation and stochastic gradient descent to minimise prediction error and refine feature representations relevant to diagnostic classification. The high-resolution images were divided into smaller patches, enabling the model to assign probabilities to diagnostic labels at both patch and whole-slide levels. It achieved AUCs of 0.99 for invasive carcinoma and 0.98 for DCIS across a diverse validation cohort, showing near-human-level accuracy12. Notably, the AI model served as a second-read system to detect misdiagnosed cases, reinforcing its value as a real-time quality control tool. This study exemplifies how well-curated datasets and granular annotations can result in high-performing diagnostic AI models.
In contrast, Malherbe emphasises the underexplored potential of AI in decoding tumour microenvironment interactions, especially the spatial relationships between immune, stromal, and tumour cell populations13. Despite the increasing availability of multiplex IHC and spatial omics technologies, most AI models reduce this complexity into simplified feature vectors or overlook spatial context altogether. For example, models processing multiplexed ion beam imaging (MIBI) and co-detection by indexing (CODEX) data often reduce the spatial organisation of immune and stromal cells into scalar quantities which is single summary values that condense rich, multi-dimensional spatial data into basic metrics, such as the total number of immune cells, the average distance between cells, or the proportion of a specific cell type. These scalar values discard spatial topology, meaning that although the presence of individual cells is captured, the biologically meaningful spatial interactions such as immune cells clustering around tumour cells, which may indicate an active immune response will be lost14,15. This simplification can obscure key spatial patterns that are highly relevant for predicting or interpreting immunotherapy outcomes13. While Malherbe highlighted this limitation and recommended the use of graph neural networks (GNNs) and spatially resolved transcriptomics as possible avenues, these strategies were not formalised into a structured model. Building upon this insight, the present review proposes a conceptual framework (Figure 2) that explicitly integrates GNNs with spatially resolved omics data to capture the biological relevance of cell-cell interactions in breast cancer. This model illustrates how GNNs can preserve spatial topology while incorporating domain-specific biological priors, thereby addressing the limitations of conventional CNN-based models that flatten tissue architecture into simplified statistics. By advancing Malherbe’s suggestion into a schematic pipeline, our framework provides a visual roadmap for how AI could be embedded into immuno-oncology workflows to enhance biomarker interpretation and immunotherapy decision-making.
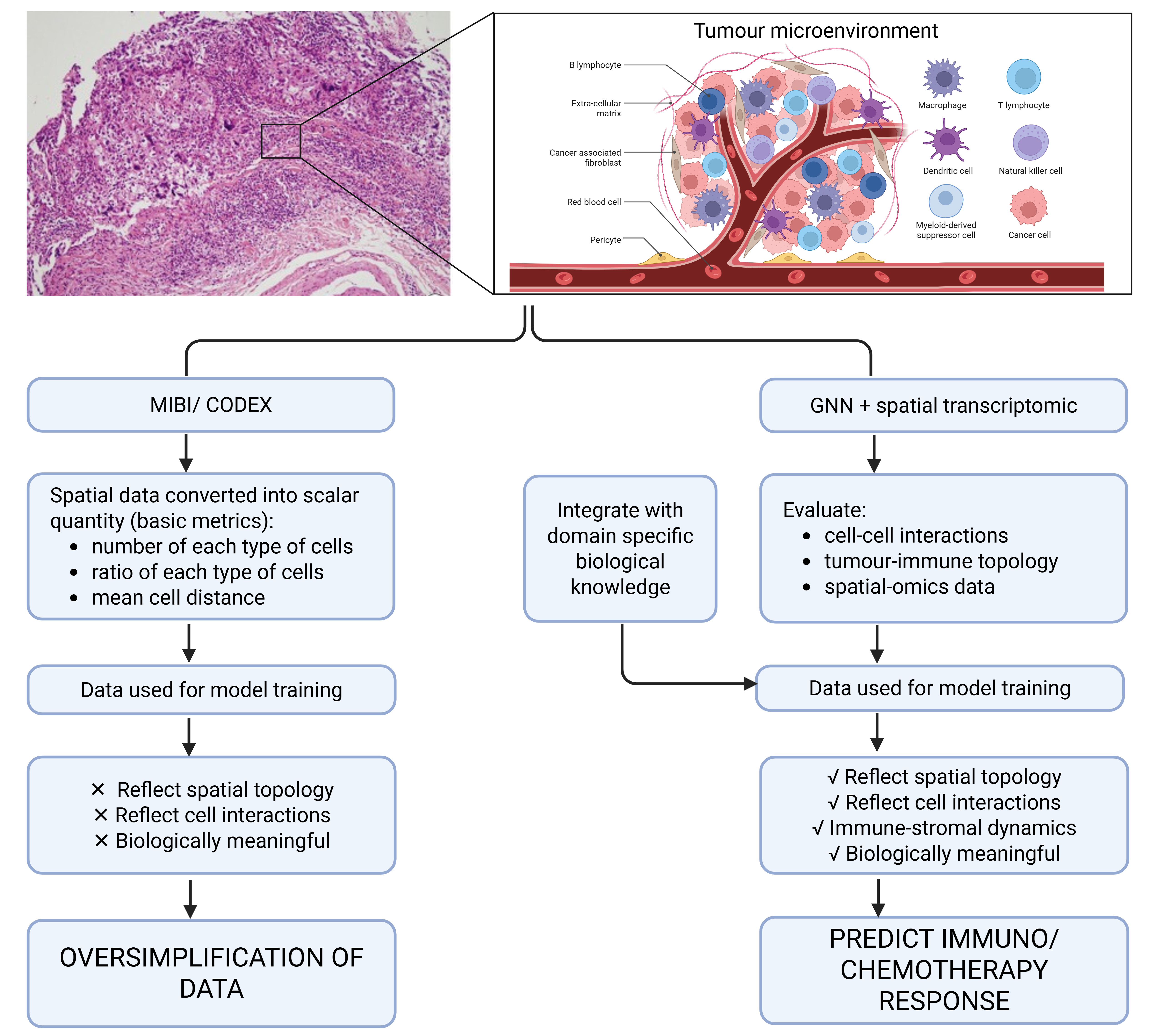
Conceptual framework proposed in this review: an AI-based model integrating GNNs with spatially resolved transcriptomics to capture tumour-immune spatial interactions. This schematic was created de novo in BioRender and does not reproduce or adapt any previously published figure. It builds upon the limitations highlighted by Malherbe (2021) but represents a novel conceptual model developed for this review.
Choudhury & Perumalla (2021) developed a CNN-based model to classify breast cancer using 50x50 pixel image patches derived from digitised H&E-stained histopathology slides, labelled as invasive ductal carcinoma (IDC+) or non-IDC (IDC-)16. The model architecture included convolution and pooling layers, fully connected layers, and a softmax output for probability generation. Given the dataset’s small size of 5,547 patches, the authors used data augmentation such as image rotation to improve generalisability16. While the model achieved a classification accuracy of 78.4%, it lacked external validation and failed to benchmark against human pathologists. Tumour biomarker integration was also absent. These limitations reduce the model’s applicability in clinical workflows. Compared to Huang’s or Sandbank’s studies, this Choudhury & Perumalla highlighted how dataset size and lack of external benchmarks can constrain AI model performance and translational impact.
AI-based prediction of tumour biomarkers expression and treatment outcome in breast cancer patients using histopathology data
The study by Meshoul et al. (2022) advances the field by predicting biomarker status of ER, PR, and HER2 directly from H&E-stained images, bypassing the need for IHC staining17. By training deep learning models on thousands of H&E-stained slides with biomarker labels verified by corresponding IHC results, the authors developed an interpretable AI framework capable of predicting ER, PR, and HER2 expression. The model employed a CNN trained in a supervised manner, in which image patches were paired with known expression levels and optimised through backpropagation. Through backpropagation, the network learned to identify morphological features such as nuclear size, chromatin texture, and glandular structure that statistically correlated with specific biomarker profiles. Although the model does not detect antigen presence via biochemical interaction, it functions as a surrogate by inferring expression likelihood based solely on visual morphological cues, thus effectively simulating IHC status prediction through image-based inference17. This approach not only reduces reagent use and turnaround time but also demonstrates the feasibility of AI-based histology-to-phenotype prediction as a potential alternative to conventional biomarker assays.
Gamble et al. (2021) extended this approach by applying AI to infer HER2, ER, and PR biomarker status directly from histomorphological features, thereby supporting the emerging concept of digital-only biopsy analysis18. Their CNN-based model was trained using annotated H&E-stained slides from The Cancer Genome Atlas Breast Invasive Carcinoma (TCGA-BRCA) cohort, with corresponding IHC data serving as the reference standard for biomarker status. The model learned to associate specific histological patterns such as glandular architecture, nuclear pleomorphism, and overall tissue organisation with known biomarker expression profiles. Although the model did not rely on biochemical detection, it effectively simulated IHC scoring by leveraging statistically derived visual cues, thereby offering a cost-effective and scalable strategy for large-scale biomarker profiling. The model achieved AUC values of 0.91 for ER, 0.95 for PR, and 0.93 for HER2 prediction, indicating high discriminative performance. The AUC metric, or area under the receiver operating characteristic (ROC) curve, reflects the model’s ability to distinguish between positive and negative cases across all possible decision thresholds. An AUC of 1.0 denotes perfect classification, while 0.5 indicates performance no better than random guessing18. Thus, the AUC values reported by Gamble et al. suggest that the model achieved excellent discriminative power in classifying biomarker expression status from histomorphological features alone. However, the study did not report sensitivity or specificity metrics, nor did it include a direct comparison between model predictions and conventional IHC scoring. Furthermore, the model’s generalisability was constrained by its reliance on a single dataset (TCGA-BRCA), and external validation across diverse clinical settings was not conducted.
Dodington et al. (2021) demonstrated the capability of AI to evaluate both treatment response and molecular subtype prognosis across distinct categories of breast cancer, including luminal A, luminal B, HER2-enriched, basal-like, and triple-negative subtypes19. Each of these molecular classifications carries specific clinical implications: luminal A tumours are typically hormone receptor-positive with low proliferative activity and favourable prognosis; luminal B tumours exhibit higher proliferation and may require chemotherapy; HER2-enriched tumours overexpress HER2 and benefit from targeted therapies; and basal-like and triple-negative subtypes tend to be more aggressive, lacking hormone receptor and HER2 expression, often resulting in poorer outcomes with limited therapeutic options2. To address this clinical heterogeneity, Dodington and colleagues developed a supervised machine learning model trained on nuclear morphometric features such as nuclear size, shape, and chromatin texture that were extracted from digitised H&E-stained biopsy slides19. Crucially, these slides were labelled with patient-level therapy outcomes, enabling the model to learn associations between pre-treatment histomorphology and pathological complete response to neoadjuvant chemotherapy. The model achieved an F1 score of 0.87, indicating a strong balance between sensitivity and precision in identifying responders while minimising false predictions19. Pathological complete response is defined as the absence of residual invasive cancer in the breast and axillary lymph nodes following treatment (ypT0/is ypN0) and is widely recognised as a surrogate marker for improved long-term survival in aggressive breast cancer subtypes20. This performance significantly outperformed a logistic regression baseline, suggesting the model was better equipped to detect subtle morphological patterns predictive of treatment efficacy. Notably, their analysis revealed that specific nuclear morphologies correlate with therapeutic response, supporting the hypothesis that tumour architecture and cellular characteristics encode predictive cues. However, the study’s limitations include its relatively small retrospective cohort (n = 82), risk of selection bias, and the absence of external validation across multiple institutions.
Building on these developments, Garberis et al. (2025) introduced a deep learning model known as RlapsRisk, designed to predict five-year metastasis-free survival (MFS) in patients with ER+/HER2- early breast cancer using routine HES-stained slides21. The model was trained on a large retrospective cohort (GrandTMA; n = 1429) and externally validated on the prospective CANTO cohort (n = 889). It utilised a multi-stage architecture that incorporated self-supervised learning for feature extraction and a multiple instance learning (MIL) approach to predict outcomes from entire whole-slide images. Notably, RlapsRisk captured prognostic morphological patterns such as high nuclear pleomorphism, mitotic activity, and tumour-stroma spatial interactions without relying on manual annotations by pathologists. The resulting risk score demonstrated independent prognostic value beyond established clinico-pathologic factors and improved discrimination performance when combined with standard clinical models (C-index improved from 0.76 to 0.80 in the validation cohort, p < 0.005)21. Importantly, the model showed robust stratification performance across clinically relevant subgroups including patients with intermediate histological grade or clinical risk and provided interpretable outputs via tile-level Shapley value analysis, supporting clinical trust and transparency. By directly addressing the clinical endpoint of distant recurrence and requiring only standard digital slides, this study exemplifies the real-world potential of AI to enhance prognostic assessment and inform adjuvant treatment decisions, particularly where access to genomic assays remains limited.
Tanveer et al. (2025), Jones et al. (2022), Choudhury & Perumalla (2021), and Nassif et al. (2022) collectively reinforce the central role of CNNs in breast cancer diagnostics, particularly in image-based classification and feature extraction16,22,23,24. However, these studies differ substantially in methodological rigor, dataset scale, and clinical applicability. Some rely on small, retrospective cohorts lacking external validation, while others incorporate more diverse data sources yet fall short in explainability or clinical workflow integration. For instance, Tanveer et al. developed an ensemble machine learning framework that combined radiomic features derived from breast imaging with serum protein biomarker data. The model was trained using a supervised learning strategy in which multimodal features were input into several base classifiers such as support vector machines and random forests and subsequently aggregated by a meta-classifier to improve prediction robustness. Although the model reported an accuracy of 91.2% in cancer detection, it was not externally validated on independent or multicentre datasets, thereby limiting its generalisability22.
Jones proposed a hybrid deep learning pipeline that integrated imaging and clinical metadata, including age, menopausal status, family history, hormone receptor status, tumour size, and comorbidities23. Their fusion architecture combined features extracted from digital mammography with those from electronic health records, using convolutional and fully connected layers. The model was trained on labelled diagnostic outcomes to predict both malignancy risk and molecular subtype23. However, the study omitted key components related to model interpretability such as feature attribution methods or saliency mapping and have not address data governance issues, including consent, data-sharing policies, and deployment logistics in clinical settings.
Taken together, these studies emphasise the technical promise of CNNs but also reveal recurring limitations such as insufficient explainability, lack of prospective or external validation, and limited integration into real-world diagnostic workflows. Critically, few of the models have progressed beyond in silico evaluation to full clinical deployment. As such, their robustness in operational environments where variability in patient demographics, imaging protocols, and institutional infrastructure inevitably remains untested.
Scoping synthesis of findings by data modality, validation design and explainability.
Across the eleven reviewed studies, three major thematic patterns emerge that collectively define the current state of AI applications for tumor biomarker detection in breast cancer: (1) data modality and diagnostic scope, (2) validation and generalisability, and (3) explainability and clinical integration (Table 2).
Data modality and diagnostic scope.
AI has demonstrated remarkable adaptability across diverse diagnostic modalities, encompassing serum protein markers, histopathology, and predictive modelling of biomarker expression based on morphology. Serum-based frameworks such as OncoSeek (Mao et al., 2023; Luan et al., 2023) leverage ensemble learning across multiple circulating proteins, effectively overcoming the high false-positive rates typical of fixed cutoff methods used in conventional assays like CA15-3 and CEA. In contrast, histopathology-driven approaches, including those by Sandbank et al. (2022), Huang et al. (2023), and Gamble et al. (2021), rely on CNNs to extract complex spatial features from digitised H&E or HES-stained slides, achieving near-human classification accuracy. Meanwhile, integrative or multimodal frameworks remain scarce; only Jones et al. (2022) and Tanveer et al. (2025) have attempted to combine imaging, serum, or clinical metadata within a unified pipeline. This trend toward unimodal development reflects a persistent disconnection between laboratory assays and clinical imaging workflows. Bridging this gap through true multimodal architectures, capable of integrating IHC, genomic, radiomic, and serum data, represents a key frontier for clinically relevant AI models.
Validation and generalisability.
Model validation emerged as the most critical methodological bottleneck across the field. While large-scale external validation was achieved in only a few cases, most notably Garberis et al. (2025), Luan et al. (2023), and Sandbank et al. (2022), the majority of studies relied on internal or single-centre datasets, often retrospective and underpowered. This narrow validation base raising concerns about generalisability across varied laboratory contexts. For instance, models trained exclusively on Western or urban cohorts may perform poorly when deployed in underrepresented or resource-limited settings, where staining quality, instrumentation, or population demographics differ substantially. Moreover, inconsistent performance reporting, where most studies reported AUCs but few included sensitivity, specificity, or calibration will further complicates comparison and reproducibility.
Despite the dominance of CNN architectures, performance varied substantially even for similar diagnostic tasks. For example, while Gamble et al. (2021) achieved AUCs above 0.9 for ER/PR/HER2 prediction, other histopathology-based studies such as Huang et al. (2023) and Dodington et al. (2021) reported lower or incomplete metrics. This heterogeneity likely reflects differences in dataset size, staining protocols, and absence of harmonised benchmarking. Future research should prioritise multi-institutional validation and harmonised benchmarking to ensure reproducibility.
Explainability and clinical integration.
A further recurring theme is the limited interpretability of deep learning models. With the exception of Meshoul et al. (2022), who applied SHAP-based interpretation, most CNN-based systems operate as “black boxes,” generating predictions without transparent biological or morphological justification. This opacity constrains clinician trust and hinders regulatory approval. Although high AUCs (e.g., 0.91–0.95 for ER/PR/HER2 prediction in Gamble et al., 2021) demonstrate technical performance, the lack of explainable AI mechanisms, such as saliency or attention mapping, continues to impede clinical adoption despite high reported AUCs. Few models have progressed beyond retrospective validation to deployment in pathology or diagnostic laboratories. Regulatory and interoperability barriers further delay translation. Moving forward, embedding saliency mapping, attention-based visualisation, and human-AI collaborative validation into model design could facilitate both interpretability and trustworthiness. Beyond algorithmic transparency, integrating health-economic and human-factor analyses into AI implementation research is essential for sustainable clinical adoption (Table 2).
Together, these thematic insights reveal a dynamic yet fragmented research landscape. The reviewed studies nonetheless highlight the growing potential of AI to transform tumour-biomarker detection for better breast-cancer diagnostics; however, several limitations must be addressed to ensure clinical relevance and long-term integration (Figure 3).
Thematic synthesis of AI applications based on data modality, validation design and explainability.
| Theme | Key Patterns (Across 11 Studies) | Representative Examples | Main Gaps / Needs |
|---|---|---|---|
| Data Modality & Diagnostic Scope | AI used mainly in serum (OncoSeek), histopathology (CNN-based H&E/HES models) or morphology-to-biomarker prediction; very few true multimodal designs. | Mao 2023; Luan 2023; Sandbank 2022; Gamble 2021; Jones 2022. | Integration across imaging + omics + clinical data remains rare; need for unified multimodal pipelines. |
| Validation & Generalisability | Most studies internal, single-centre, retrospective; only 3/11 had external or multicentre validation. | Garberis 2025; Luan 2023; Sandbank 2022. | Require large, prospective, multi-institutional validation and consistent reporting of sensitivity, specificity, calibration. |
| Explainability & Clinical Integration | Only Meshoul 2022 used formal XAI (SHAP); others remain “black-box.” Few clinical deployments. | Meshoul 2022; Gamble 2021; Choudhury 2021. | Need embedded saliency/attention mapping, human-AI review, and integration with regulatory, cost-effectiveness, and workflow studies. |
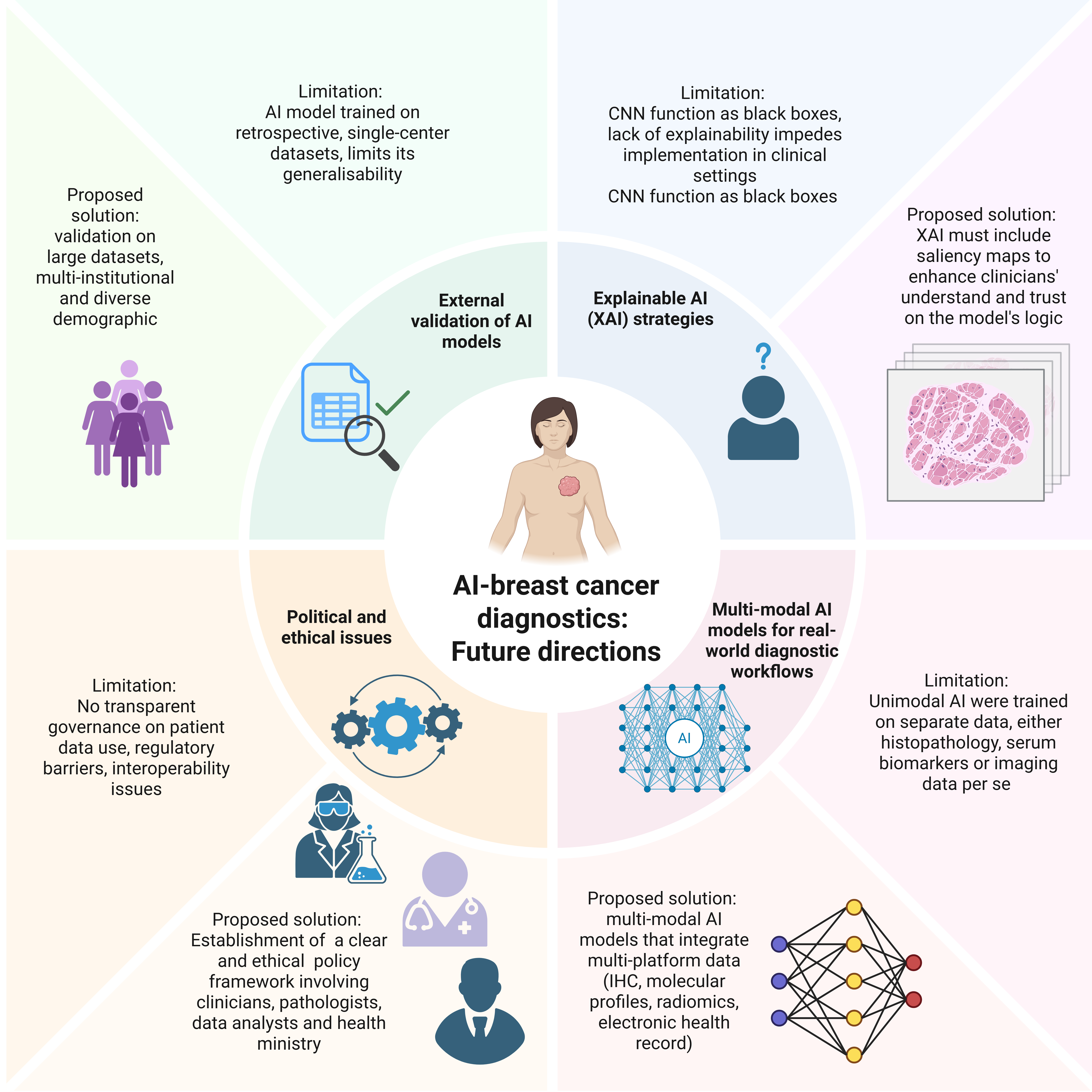
Limitations in current AI use in breast cancer diagnosis and the proposed solutions for future directions. The schematic summarises recurrent challenges identified across the reviewed studies, including limited external and prospective validation, reliance on unimodal data sources, insufficient model explainability, and barriers to clinical integration. Future directions include multi-institutional validation, development of multimodal AI frameworks, integration of XAI strategies, and consideration of regulatory and implementation requirements. Created with BioRender.com.
Limitations and Future Directions
The reviewed studies highlight the growing potential of AI to transform tumour biomarker detection for better breast cancer diagnostics. However, several limitations emerge that must be addressed to ensure clinical relevance, fairness, and long-term integration (Figure 3). Consistent with the findings above, external validation remains scarce. Most models were trained on retrospective, single-centre datasets, such as those in studies by Mao, Tanveer, and Dodington, which raise generalisability concerns. In addition, the issue of representation remains a significant ethical and clinical concern. Many AI models are trained on data from high-resource, Western settings, which may lead to performance disparities when deployed in more diverse or resource-constrained populations. Future research should include prospective, registry-based external validation of RlapsRisk in Asian and resource-limited cohorts to confirm generalisability, as well as randomised controlled trials comparing OncoSeek triage performance against conventional CA15-3 and CEA testing. More broadly, studies should prioritise validation on large, multi-institutional, and demographically diverse datasets to ensure robustness across varied clinical environments. Furthermore, model development should explicitly include underrepresented groups to mitigate bias and promote equitable care.
Another critical issue is the lack of explainability in many high-performing models. Deep learning systems, particularly CNNs employed by Choudhury, Gamble, and Meshoul, often function as black boxes that generate predictions without transparent reasoning or feature attribution. For broader clinical adoption, XAI strategies must be integrated, including saliency maps and attention mechanisms that enable clinicians to interpret the model’s decision-making process. Moreover, morphology-to-biomarker inference models, such as those proposed by Meshoul and Gamble, often lack biologically grounded explanations linking tissue architecture to protein expression. Although these models can predict ER, PR, or HER2 status from visual cues in H&E-stained slides, future work should aim to elucidate the mechanistic associations between morphology and molecular phenotype to enhance interpretability and scientific robustness. To advance clinical readiness, future studies should conduct prospective human-AI collaborative evaluations, for instance, by testing XAI-enhanced digital pathology systems in real-world pathology workflows. Pilot deployment of interpretable CNN frameworks within hospital laboratories, paired with clinician-in-the-loop validation, would provide essential evidence for regulatory approval and workflow optimisation.
Most reviewed models also rely on a unimodal architecture, using only a single data modality such as histopathology, serum markers, or imaging data. This narrow focus limits their ability to capture the multifaceted nature of cancer diagnostics. In response to this limitation, Jones and Malherbe advocate for the development of multimodal AI models that integrate diverse diagnostic data streams including IHC, molecular assays, radiomics, and electronic health records to more accurately reflect real-world clinical workflows and improve predictive performance. In the future, prospective feasibility studies combining H&E-derived morphological features with serum-based biomarker panels, such as OncoSeek, could evaluate whether integrated models outperform unimodal systems in terms of sensitivity, specificity, and interpretability. Similarly, cross-platform projects linking histopathology-derived deep features with transcriptomic or proteomic data could elucidate tumour biology, thus enhancing AI predictions and improving their translational relevance.
Despite promising diagnostic performance, the real-world implementation of AI tools remains limited. While models developed by Sandbank and Luan exhibit high technical accuracy, few have achieved clinical integration due to systemic challenges, including regulatory barriers, interoperability limitations, and the absence of comprehensive cost-effectiveness analyses. To assess feasibility and impact, collaborative efforts involving health economists, clinicians, and IT professionals are essential.
Ethical and political considerations demand urgent attention, particularly in relation to data ownership, patient privacy, and equitable benefit sharing. AI systems in oncology are frequently developed using large patient datasets, yet transparent governance and fair return of value are often lacking. Recent regulatory frameworks provide important direction as in the EU Artificial Intelligence Act (2024) which formally classifies diagnostic AI systems as “high-risk,” requiring transparency, traceability, and human oversight throughout development and deployment25. In parallel, the FDA’s released guidance on Artificial Intelligence/Machine Learning-Enabled Software as a Medical Device (AI/ML-based SaMD) highlights Good Machine Learning Practices (GMLP), continuous monitoring, and lifecycle accountability to ensure patient safety and trust26. These initiatives underscore the need for developers and clinicians to align AI-driven biomarker detection tools with evolving international standards. Embedding such principles into research pipelines is critical not only for legal compliance but also for fostering public trust, ensuring informed consent, and promoting fair distribution of benefits arising from AI-based diagnostics27,28.
Finally, although this review is focused on AI models for tumour biomarker detection, future syntheses may benefit from explicitly examining multimodal pipelines such as radiomics combined with serum and histopathology, and mapping evidence gaps against specific steps of the clinical workflow. Such analyses could further clarify how AI tools might be embedded into end-to-end patient management pathways.
Conclusion
Taken together, the reviewed evidence highlights both the promise and the limitations of current AI applications in breast cancer biomarker detection. Four central insights emerge. First, histopathology-based AI continues to dominate the evidence base, while genuine multimodal integration, which combines histology, serum, imaging, and molecular data, remains uncommon despite its clinical importance. Second, explainability is the weakest methodological link, with very few studies incorporating interpretable tools, despite their essential role in building clinician trust. Third, validation rigour remains uneven: a minority of externally validated studies provide robust and reproducible evidence, whereas most remain exploratory or rely solely on internal validation. Finally, while reported performance metrics are encouraging, they are often inconsistent and rarely benchmarked against clinical gold standards such as IHC or long-term survival outcomes.
Addressing these methodological and translational challenges is critical. The long-term success of AI in this domain depends on resolving issues related to validation, explainability, clinical workflow integration, inclusivity for diverse populations, and ethical governance surrounding data use. Future research should prioritise prospective multicentre validation, transparent and explainable modelling, multimodal data integration, and harmonised reporting standards. Only by addressing these limitations can AI systems transition from experimental proof-of-concept models to clinically trustworthy tools embedded within routine breast cancer diagnostics and treatment decision-making.
Abbreviations
AI: Artificial intelligence, AR: Androgen receptor, AUC: Area under the curve, BRCA1/2: Breast cancer gene 1 and 2, CA 15-3: Cancer antigen 15-3, CA 27.29: Cancer antigen 27.29, CEA: Carcinoembryonic antigen, CNN: Convolutional neural network, DCIS: Ductal carcinoma in situ, ER: Estrogen receptor, GNN: Graph neural network, H&E: Hematoxylin and eosin, HER2: Human epidermal growth factor receptor 2, HES: Hematoxylin-eosin-saffron, IHC: Immunohistochemistry, IDC: Invasive ductal carcinoma, Ki-67: antigen Kiel 67, MIL: Multiple instance learning, MFS: Metastasis-free survival, PR: Progesterone receptor, ROC: Receiver operating characteristic, SOP: Standard operating procedure, TCGA-BRCA: The Cancer Genome Atlas Breast Invasive Carcinoma, WSI: Whole slide image, XAI: Explainable artificial intelligence.
Acknowledgments
We would like to thank Universiti Teknologi MARA and Ministry of Higher Education Malaysia for funding this research through Fundamental Research Grant Scheme (DP KPT FRGS/1/2019/SKK15/UITM/03/1), Project ID 15531, RMC file number 600-IRMI/FRGS5/3 (286/2019).
Author’s contributions
Design & Conceptualisation, Investigation, Writing - Review & Editing (SFM); Writing - Original draft, table & figure (NAA, NFM); Writing - Original draft (MNFR, SMHD, NAR). All authors read and approved the final manuscript.
Funding
Fundamental Research Grant Scheme (DP KPT FRGS/1/2019/SKK15/UITM/03/1), Project ID 15531, RMC file number 600-IRMI/FRGS5/3 (286/2019).
Availability of data and materials
Data and materials used and/or analysed during the current study are available from the corresponding author on reasonable request.
Ethics approval and consent to participate
Not applicable.
Consent for publication
Not applicable.
Declaration of generative AI and AI-assisted technologies in the writing process
The authors declare that they have not used generative AI (a type of artificial intelligence technology that can produce various types of content including text, imagery, audio and synthetic data.
Competing interests
The authors declare that they have no competing interests.

